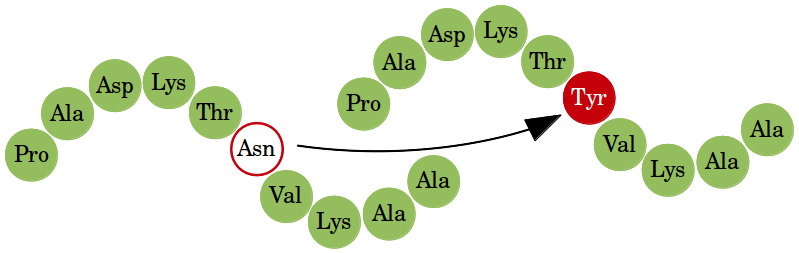
Prediction of mutation-induced stability changes in protein variants with EASE programs, data sets, and other resources
Contact authors
* Lukas Folkman
* please report any bugs or problems
e-mail: 
NEW: EASE-MM predictions are now available as a public web-server.
EASE-MM: Evolutionary, Amino acid, and Structural Encodings with Multiple Models
The analysis of the results from our previous work (EASE-AA, see below) revealed that the classification performance varies considerably for the exposed and buried residues. Also, mutations in coil residues seemed to be harder to classify than mutations in helical and sheet residues. Therefore, we proposed a method which combines feature-based multiple models with each model specifically designed for different types of mutations. The new method (EASE-MM) not only outperformed our previous work (EASE-AA) but yielded a more balanced performance for different types of mutations (buried and exposed; helix, sheet, and coil; small and large stability changes). This work was presented at the ISMB/ECCB Single Nucleotide Polymorphism Special Interest Group (SNP-SIG 2013) meeting in Berlin.
* data sets
* web-server
how to cite (the web-server and real-value prediction):
Folkman, L., Stantic, B., Sattar, A.& Zhou, Y. (2016) ‘EASE-MM: sequence-based prediction of mutation-induced stability changes with feature-based multiple models’, Journal of Molecular Biology, doi:10.1016/j.jmb.2016.01.012.
how to cite (two-state classification):
Folkman, L., Stantic, B. & Sattar, A. (2014), ‘Feature-based multiple models improve classification of mutation-induced stability changes’, BMC Genomics
15(Suppl 4), S6. [pdf] [supplements] [publisher] [copyright]
EASE-AA: Evolutionary And Structural Encodings with Amino Acid parameters In this work, we addressed the problem of the overestimation of the prediction performance of currently available methods. We introduced an evaluation scheme which provides a realistic estimate of prediction performance for previously unseen non-homologous proteins. We compiled a dataset of 1,914 mutations from the public ProTherm database. Out of the 1,914 mutations, we had to fix more than 230 erroneous entries. Finally, we proposed a method (EASE-AA) which considerably outperformed our previous work (Folkman et al. 2013) and other available methods. Our results were presented at the Asia Pacific Bioinformatics Conference (APBC 2014) in Shanghai. * data sets * source code * installation instructions how to cite: Folkman, L., Stantic, B. & Sattar, A. (2014), ‘Towards sequence-based prediction of mutation-induced stability changes in unseen non-homologous proteins’, BMC Genomics 15(Suppl 1), S4. [pdf] [publisher] [PubMed] [copyright]